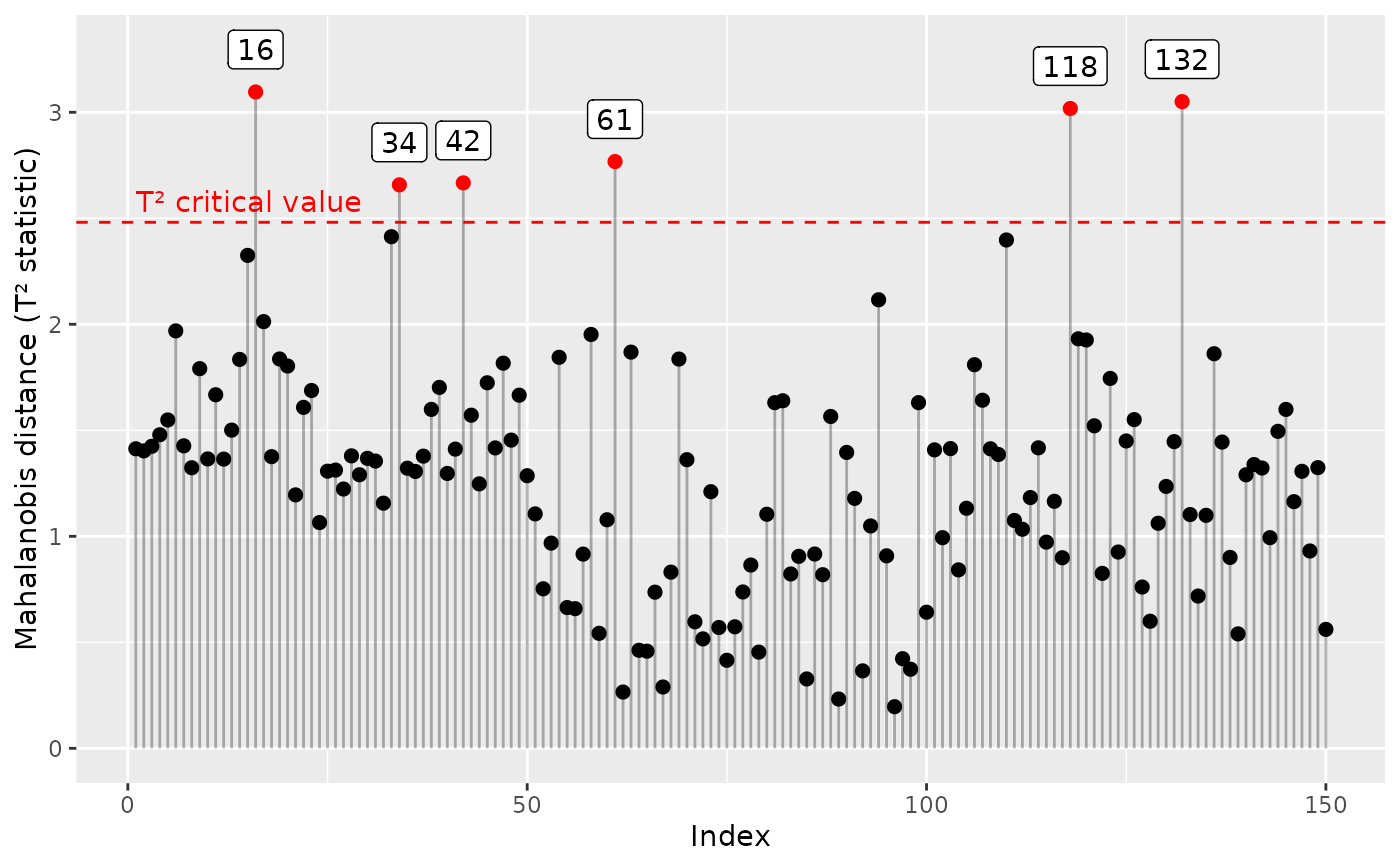
Calculate the T2 statistic for individual points
outliers.RdCalculate the T2 statistic or Mahalanobis distance for individual points
Arguments
- x
A matrix or data frame with two columns
- level
Either coverage probability (for type = "t2data" or "c2data") or confidence level (for type = "t2mean").
- robust
If TRUE, then robust estimates of mean and covariance are used
- type
what type of statistic should be calculated; can be t2data (for data coverage), t2mean (for difference from a mean) or "c2data" (for coverage calculated with the chi squared statistic)
- labels
Optional labels to use on the plot instead of rownames
Value
A data frame with one row per point including the columns d2 (squared mahalanobis distance) t2crit (critical T squared value for the given level), c2crit (critical X squared value for the given level) and is_outlier (logical, whether d2 > t2crit or d2 > c2crit, depending on type).
Details
The function can use a robust estimator of location and scatter
using the covMcd function, which uses the
Maximum Covariance Determinant (MCD) estimator. Note that while this
results in ellipses which are more resistent to outliers, the
interpretation slightly changes, as the T2 statistic used is only an
approximation in this case. In other words, use it for visualisation and
QC, but not for statistical testing.
See also
hotelling_ellipse for more information on the
differences between t2data, t2mean and c2data modes.